 |
 |
 |
 |
 |
 |


Dr. Dalia Barsyte-Lovejoy, PhD is an Assistant Professor at the Department of Pharmacology and Toxicology, UofT, and Principal Investigator at the SGC-Toronto working to understand fundamental regulatory mechanisms of epigenetic proteins and their pharmacological modulation in cancer. Her research focuses on chemical probe discovery, disease mechanisms, and therapeutic target validation. Her research at the SGC-Toronto has played an instrumental role in the cellular characterization of chemical probes and functional target investigations. Importantly, her chemical probe work provided 25 extensively characterized compounds that have helped shape the emerging field of epigenetics and enabled over 50 collaborative projects that are uncovering new epigenetic mechanisms in cancer and its treatment.
We are interested in understanding the mechanism of epigenetic regulators that control cell growth, differentiation, carcinogenesis and adaptation. Through use of the chemical probes to target the epigenetic regulators we are investigating the cellular functions of these proteins and their potential to become therapeutic targets in such diseases as cancer. Our work focuses on protein lysine and arginine methyltransferases involved in transcription, genome stability, RNA metabolism and variety of cell processes determined by the substrates of these cellular enzymes. The PRMT, arginine methyltransferase, family has especially wide scope of substrates ranging from histones to signaling molecules, enzymes and structural proteins. Through multidisciplinary research that includes cell and chemical biology, protein structural biology and many collaborative studies with colleagues across industry and academia, the SGC chemical probes project has generated several probes for various PRMTs. We are currently using the chemical probes to explore the cellular pathways that are controlled by PRMT enzymes in especially poor prognosis acute myeloid leukemia patient cells. The long term patient leukemia cell cultures developed in our group enable us to investigate the cancer cell growth, drug resistance and self-renewal capacity.
Epigenetics and chromatin architecture regulators
| Epigenetics and chromatin architecture regulators | |
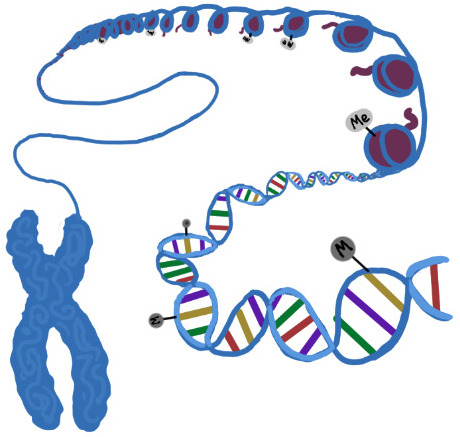 |
Epigenetics is about how the DNA code is regulated. Proteins that bind/modify DNA and histones play essential roles in cell identity determination, transcription, and genome maintenance. They are often responsible for diseases such as cancer or uncontrolled inflammation. We are studying how epigenetic proteins regulate normal cell processes and how these are subverted in disease. |
|
Chemical probes as tools for cancer target discovery |
|
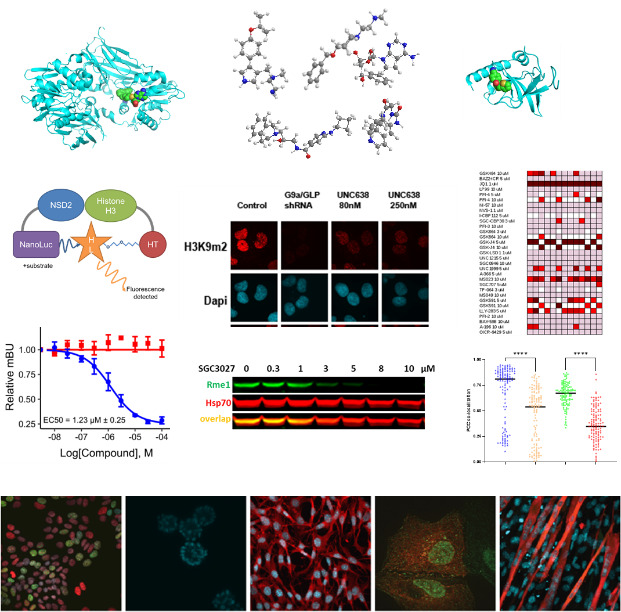 |
To study epigenetic modifier proteins, we need genetic and pharmacological tools. Chemical probe compounds that potently and selectively inhibit or degrade the target proteins in cells provide tools for modulating activating/repressing histone marks and other cellular signaling pathways. By discovering and using chemical probes, we expand our understanding of the protein function and its therapeutic utility to establish a biological rationale in cancer therapy.
For more information please see Group Research and Members tab above. |
Link to Open Lab notebooks that features science community posts on our various projects https://openlabnotebooks.org/
Nat. Chem. Biol.. 2017 . doi: 10.1038/nchembio.2306
PMID: 28135237
Nat. Chem. Biol.. 2017 . doi: 10.1038/nchembio.2282
PMID: 28114273
J. Med. Chem.. 2016 . doi: 10.1021/acs.jmedchem.6b01033
PMID: 27584694
J. Proteome Res.. 2016 . doi: 10.1021/acs.jproteome.6b00220
PMID: 27163177
Meth. Enzymol.. 2016 574:79-103. doi: 10.1016/bs.mie.2016.01.011
PMID: 27423858
ACS Chem. Biol.. 2015 . doi: 10.1021/acschembio.5b00839
PMID: 26598975
Nature. 2015 525(7568):206-11. doi: 10.1038/nature15251
PMID: 26331536
Nat. Chem. Biol.. 2015 . doi: 10.1038/nchembio.1859
PMID: 26167872
Angew. Chem. Int. Ed. Engl.. 2015 . doi: 10.1002/anie.201412154
PMID: 25728001
Proc. Natl. Acad. Sci. U.S.A.. 2014 . doi: 10.1073/pnas.1407358111
PMID: 25136132
ACS Chem. Biol.. 2013 . doi: 10.1021/cb400133j
PMID: 23614352
Nat. Chem. Biol.. 2013 . doi: 10.1038/nchembio.1157
PMID: 23292653
Biochem. J.. 2013 449(1):151-9. doi: 10.1042/BJ20121280
PMID: 22989411
Nat Commun. 2012 3:1288. doi: 10.1038/ncomms2304
PMID: 23250418
Cell. 2012 149(1):214-31. doi: 10.1016/j.cell.2012.02.013
PMID: 22464331
Nat. Chem. Biol.. 2011 7(8):566-74. doi: 10.1038/nchembio.599
PMID: 21743462
Nat Rev Drug Discov. 2021 . doi: 10.1038/s41573-021-00159-8
PMID: 33742187
Nat Chem Biol. 2021 . doi: 10.1038/s41589-021-00898-0
PMID: 34782742
Nat Commun. 2020 11(1):2396. doi: 10.1038/s41467-020-16271-z
PMID: 32409666
2024
Nat Chem Biol. 21.05.2024 . doi: 10.1038/s41589-024-01618-0
PMID: 38773330
Nat Commun. 01.04.2024 15(1):2809. doi: 10.1038/s41467-024-47107-9
PMID: 38561334
PNAS Nexus. 01.04.2024 3(4):pgae153. doi: 10.1093/pnasnexus/pgae153
PMID: 38665159
RSC Med Chem. 20.03.2024 15(3):1066-1071. doi: 10.1039/d3md00633f
PMID: 38516600
2023
Commun Biol. 16.12.2023 6(1):1272. doi: 10.1038/s42003-023-05655-8
PMID: 38104184
ACS Med Chem Lett. 14.12.2023 14(12):1746-1753. doi: 10.1021/acsmedchemlett.3c00386
PMID: 38116405
Exp Hematol. 10.12.2023 104135. doi: 10.1016/j.exphem.2023.11.009
PMID: 38072134
Bioorg Med Chem Lett. 26.09.2023 129488. doi: 10.1016/j.bmcl.2023.129488
PMID: 37770003
Methods Mol Biol. 10.08.2023 2706:149-165. doi: 10.1007/978-1-0716-3397-7_11
PMID: 37558947
Methods Mol Biol. 10.08.2023 2706:137-148. doi: 10.1007/978-1-0716-3397-7_10
PMID: 37558946
J Med Chem. 27.07.2023 . doi: 10.1021/acs.jmedchem.3c00314
PMID: 37499118
J Am Chem Soc. 28.03.2023 . doi: 10.1021/jacs.3c01421
PMID: 36976643
J Med Chem. 22.03.2023 . doi: 10.1021/acs.jmedchem.2c02132
PMID: 36948210
Nat Commun. 13.03.2023 14(1):1364. doi: 10.1038/s41467-023-36829-x
PMID: 36914634
J Med Chem. 20.02.2023 . doi: 10.1021/acs.jmedchem.2c01933
PMID: 36802665
2022
Front Physiol. 17.12.2022 13:1031264. doi: 10.3389/fphys.2022.1031264
PMID: 36523555
Biochem Biophys Rep. 06.12.2022 32:101397. doi: 10.1016/j.bbrep.2022.101397
PMID: 36467544
ACS Chem Biol. 09.09.2022 . doi: 10.1021/acschembio.2c00451
PMID: 36084291
Biochim Biophys Acta Gene Regul Mech. 27.07.2022 194845. doi: 10.1016/j.bbagrm.2022.194845
PMID: 35907431
Biochim Biophys Acta Gene Regul Mech. 23.06.2022 194840. doi: 10.1016/j.bbagrm.2022.194840
PMID: 35753676
Nat Chem Biol. 16.05.2022 . doi: 10.1038/s41589-022-01024-4
PMID: 35578032
Annu Rev Biochem. 01.04.2022 . doi: 10.1146/annurev-biochem-032620-105344
PMID: 35363509
Cell Rep. 29.03.2022 38(13):110582. doi: 10.1016/j.celrep.2022.110582
PMID: 35354055
Redox Biol. 11.03.2022 51:102282. doi: 10.1016/j.redox.2022.102282
PMID: 35305370
2021
Nat Chem Biol. 15.11.2021 . doi: 10.1038/s41589-021-00898-0
PMID: 34782742
Life (Basel). 30.07.2021 11(8):. doi: 10.3390/life11080768
PMID: 34440512
ChemMedChem. 23.06.2021 . doi: 10.1002/cmdc.202100432
PMID: 34164919
SLAS Discov. 21.06.2021 24725552211019409. doi: 10.1177/24725552211019409
PMID: 34154424
J Med Chem. 13.05.2021 . doi: 10.1021/acs.jmedchem.1c00146
PMID: 33980013
Nat Rev Drug Discov. 19.03.2021 . doi: 10.1038/s41573-021-00159-8
PMID: 33742187
J Med Chem. 16.02.2021 . doi: 10.1021/acs.jmedchem.0c02160
PMID: 33591753
Nat Commun. 12.02.2021 12(1):979. doi: 10.1038/s41467-021-21204-5
PMID: 33579912
J Med Chem. 01.02.2021 . doi: 10.1021/acs.jmedchem.0c01768
PMID: 33522809
2020
Nat Commun. 14.05.2020 11(1):2396. doi: 10.1038/s41467-020-16271-z
PMID: 32409666
J. Med. Chem.. 05.05.2020 . doi: 10.1021/acs.jmedchem.0c00406
PMID: 32367723
Elife. 16.04.2020 9:. doi: 10.7554/eLife.47333
PMID: 32297856
2019
Nat Commun. 17.12.2019 10(1):5759. doi: 10.1038/s41467-019-13652-x
PMID: 31848333
Front Endocrinol (Lausanne). 30.11.2019 10:730. doi: 10.3389/fendo.2019.00730
PMID: 31781029
J. Med. Chem.. 15.08.2019 . doi: 10.1021/acs.jmedchem.9b00112
PMID: 31415173
Cancer Cell. 12.08.2019 36(2):194-209.e9. doi: 10.1016/j.ccell.2019.07.003
PMID: 31408619
Cancer Cell. 08.07.2019 36(1):51-67.e7. doi: 10.1016/j.ccell.2019.06.002
PMID: 31287992
Nat. Chem. Biol.. 08.07.2019 . doi: 10.1038/s41589-019-0310-x
PMID: 31285596
J. Med. Chem.. 01.07.2019 . doi: 10.1021/acs.jmedchem.9b00522
PMID: 31260300
Nat Commun. 29.03.2019 10(1):1436. doi: 10.1038/s41467-019-09309-4
PMID: 30926792
Front Endocrinol (Lausanne). 19.02.2019 10:22. doi: 10.3389/fendo.2019.00022
PMID: 30774623
Nat Commun. 03.01.2019 10(1):19. doi: 10.1038/s41467-018-07905-4
PMID: 30604761
2018
Nucleic Acids Res.. 20.11.2018 . doi: 10.1093/nar/gky1192
PMID: 30462309
Nucleic Acids Research. 20.11.2018 :-. doi: 10.1093/nar/gky1192
PMID: 30462309
Exp. Hematol.. 20.10.2018 . doi: 10.1016/j.exphem.2018.10.006
PMID: 30352278
ACS Med Chem Lett. 12.07.2018 9(7):612-617. doi: 10.1021/acsmedchemlett.8b00014
PMID: 30034588
J. Cell Biol.. 19.06.2018 . doi: 10.1083/jcb.201804042
PMID: 29921600
Oncotarget. 06.04.2018 9(26):18480-18493. doi: 10.18632/oncotarget.24883
PMID: 29719619
2017
J. Med. Chem.. 15.12.2017 . doi: 10.1021/acs.jmedchem.7b01674
PMID: 29244490
Nat Commun. 15.11.2017 8(1):1527. doi: 10.1038/s41467-017-01657-3
PMID: 29142305
Protein Sci.. 04.02.2017 . doi: 10.1002/pro.3129
PMID: 28160335
Nat. Chem. Biol.. 30.01.2017 . doi: 10.1038/nchembio.2306
PMID: 28135237
Nat. Chem. Biol.. 23.01.2017 . doi: 10.1038/nchembio.2282
PMID: 28114273
2016
Cancer Cell. 12.12.2016 30(6):891-908. doi: 10.1016/j.ccell.2016.11.003
PMID: 27960086
J. Med. Chem.. 01.09.2016 . doi: 10.1021/acs.jmedchem.6b01033
PMID: 27584694
J. Med. Chem.. 08.07.2016 . doi: 10.1021/acs.jmedchem.6b00668
PMID: 27390919
J. Proteome Res.. 10.05.2016 . doi: 10.1021/acs.jproteome.6b00220
PMID: 27163177
J. Med. Chem.. 13.04.2016 . doi: 10.1021/acs.jmedchem.5b01890
PMID: 27075367
Meth. Enzymol.. 16.02.2016 574:79-103. doi: 10.1016/bs.mie.2016.01.011
PMID: 27423858
J. Med. Chem.. 29.01.2016 . doi: 10.1021/acs.jmedchem.5b01772
PMID: 26824386
2015
Cancer Cell. 25.11.2015 . doi: 10.1016/j.ccell.2015.10.005
PMID: 26626085
ACS Chem. Biol.. 24.11.2015 . doi: 10.1021/acschembio.5b00839
PMID: 26598975
Nature Chemical Biology. 01.10.2015 11:815-. doi: 10.1038/nchembio1015-815b
PMID: 26379026
Nature. 10.09.2015 525(7568):206-11. doi: 10.1038/nature15251
PMID: 26331536
Nat. Chem. Biol.. 13.07.2015 . doi: 10.1038/nchembio.1859
PMID: 26167872
ACS Med Chem Lett. 09.04.2015 6(4):408-12. doi: 10.1021/ml500467h
PMID: 25893041
J. Biol. Chem.. 30.03.2015 . doi: 10.1074/jbc.M114.626861
PMID: 25825497
Angew. Chem. Int. Ed. Engl.. 27.02.2015 . doi: 10.1002/anie.201412154
PMID: 25728001
2014
Proc. Natl. Acad. Sci. U.S.A.. 18.08.2014 . doi: 10.1073/pnas.1407358111
PMID: 25136132
ChemMedChem. 17.01.2014 . doi: 10.1002/cmdc.201300450
PMID: 24443078
2013
J. Med. Chem.. 08.10.2013 . doi: 10.1021/jm401480r
PMID: 24102134
Dev. Cell. 09.07.2013 . doi: 10.1016/j.devcel.2013.05.025
PMID: 23850191
ACS Chem. Biol.. 24.04.2013 . doi: 10.1021/cb400133j
PMID: 23614352
Prog. Biophys. Mol. Biol.. 14.03.2013 . doi: 10.1016/j.pbiomolbio.2013.02.002
PMID: 23500148
Nat. Chem. Biol.. 06.01.2013 . doi: 10.1038/nchembio.1157
PMID: 23292653
Biochem. J.. 01.01.2013 449(1):151-9. doi: 10.1042/BJ20121280
PMID: 22989411
2012
Nat Commun. 18.12.2012 3:1288. doi: 10.1038/ncomms2304
PMID: 23250418
Nat. Struct. Mol. Biol.. 30.09.2012 19(11):1155-60. doi: 10.1038/nsmb.2391
PMID: 23022729
Mol. Cell. Neurosci.. 28.09.2012 52:38-50. doi: 10.1016/j.mcn.2012.09.006
PMID: 23026563
Neuroscience. 06.09.2012 219:255-70. doi: 10.1016/j.neuroscience.2012.05.069
PMID: 22698694
Cell. 30.03.2012 149(1):214-31. doi: 10.1016/j.cell.2012.02.013
PMID: 22464331
2011
J. Med. Chem.. 08.09.2011 54(17):6139-50. doi: 10.1021/jm200903z
PMID: 21780790
Nat. Chem. Biol.. 10.07.2011 7(8):566-74. doi: 10.1038/nchembio.599
PMID: 21743462
